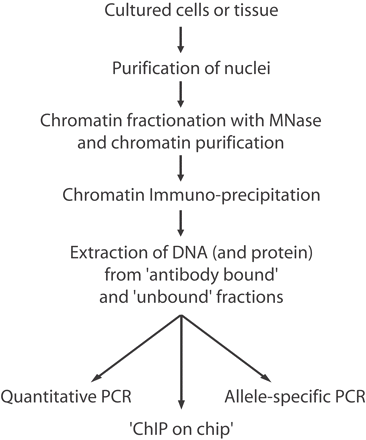
Flowchart of the procedures used to investigate site-specific covalent modifications (i.e., methylation and acetylation) on histones. In summary, nuclei are purified from fresh/frozen tissues or from cells, and the chromatin, after fractionation with micrococcal nuclease (MNase), is purified from the nuclei. This “input chromatin,” made up of fragments of up to seven nucleosomes in length, is incubated with an antiserum directed against the histone modification of interest. The antibody-bound fraction is separated from the unbound fraction and, after extraction of genomic DNA from the bound and unbound fractions, PCR technologies are applied to specifically analyze the gene or chromosomal region of interest. Precipitated DNAs can be used as probes to hybridize DNA tiling arrays (ChIP on chip) as well.










