Figure 1.
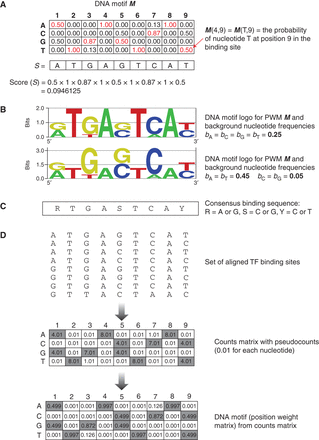
TF–DNA binding models. (A) Representation of a TF–DNA binding motif as a position weight matrix (PWM). (B) DNA motif logos generated using the enoLOGOS web server (Workman et al. 2005) for the PWM in A using a uniform background distribution bA = bC = bG = bT = 0.25 (top logo) or bA = bT = 0.45, bC = bG = 0.05 (bottom logo). (C) Consensus binding sequence derived from DNA motif in A. (D) Building a DNA motif from a set of aligned TF binding sites.








