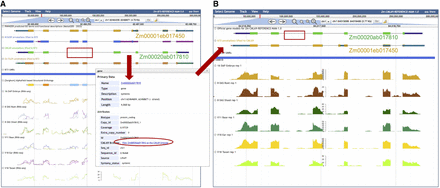
Two sample JBrowse instances in MaizeGDB. (A) JBrowse instance of the reference genome B73, version 5. The top track represents B73 gene model Zm00001eb017450. Below it are tracks of gene model annotations lifted to B73 from other genomes, including the gene model Zm00020ab017810 from the genome CML69 (green). Notice that Zm00020ab017810 is missing two exons relative to the other tracks (red rectangle). Clicking on the gene model Zm00020ab017810 opens up a pop-up box with a link (red oval) to Zm00020ab017810 in its home genome CML69 JBrowse instance. (B, top) The exons missing in the B73 JBrowse are also missing in CML69, and are missing in the B73 gene model Zm00001eb017450, which has been lifted over to CML69 because those exon sequences are not present in CML69. (Bottom) RNA-seq tracks show that there are no RNA reads associated with the missing exon space, suggesting that the exons are indeed absent and might have been deleted and are not merely the result of a misannotation.










