Metabolite Profiling of Growth Regulatory Hormones from Maize Tissue
- 1Department of Biochemistry, Center for Plant Biology, Purdue University, West Lafayette, Indiana 47907, USA
- 2Plant Genetics Research Unit, Agricultural Research Service, U.S. Department of Agriculture, Columbia, Missouri 65211, USA
- ↵3Correspondence: norman.best{at}usda.gov
Abstract
Plant regulatory small molecules, or phytohormones, are small regulatory metabolites in plants. Phytohormones regulate all aspects of plant growth and development. They include jasmonic acid, auxin, abscisic acid, salicylic acid, 1-aminocyclopropane-1-carboxylic acid, gibberellins, and brassinosteroids. Their activity is highly dependent on their concentration, and therefore accurate quantification is necessary to understand their biological role in regulating downstream targets. However, their low abundance results in low signal to noise ratios during detection. In addition, the chemical distinctions between the regulatory small molecule classes include a wide polarity range and differences in charge, which has previously prevented the simultaneous extraction and separation by chromatography of multiple regulatory small molecules. This review discusses the extraction of hormones from any maize tissue, followed by their purification and quantification, and the limitations of these approaches. Recent advancements in mass spectrometry and sample pretreatment have improved the sensitivity of techniques to accurately and simultaneously quantify multiple small regulatory plant hormones from maize tissue. These techniques should usher in a new era in which measurement of phytohormones will allow for more accurate evaluation of phytohormone roles in maize growth and development. We also highlight potential new plant regulatory hormones and discuss how the techniques described here may benefit future discovery of new classes of phytohormones.
INTRODUCTION
The control of plant growth and development is coordinated in part by the synthesis, transport, and perception of distinct small molecules collectively referred to as plant hormones. These specialized metabolites include abscisic acid (ABA), auxins, brassinosteroids (BRs), cytokinin (CK), gibberellins (GA), jasmonic acid (JA), salicylic acid (SA), and strigolactones. Hormones are specialized small signaling molecules and their signal transduction pathways and responses have distinct effects from the homeostatic signaling and growth effects of primary metabolism. Plant hormones can act distantly from their site of synthesis and responses can be developmentally regulated. Careful observation of plant phenotypic and metabolic responses requires the measurement of hormone levels in plant tissues for the interpretation of hormone-regulated phenomena. Hormones typically effect changes in plant growth and development at low concentrations. The low abundance of these metabolites in physiologically relevant situations presents the first challenge to their detection, measurement, and analysis. A second challenge has historically been the difficulty in chemically identifying the compounds, quantifying their levels, and synthesizing bona fide standards. These two challenges have often been overcome by using tissues with unusually high hormone accumulation for initial purification and compound identification. For example, BRs were first chemically identified by collecting 230 kg of Brassica napus pollen (∼2.3 × 1014 B. napus pollen grains) (Grove et al. 1979), while GAs were identified initially from Gibberella fujikuroi fungi that overproduce them (Brian et al. 1954).
Plant metabolites can be separated via liquid chromatography (LC) using a column, by reverse phase separation. A mobile phase, into which extracted metabolites are injected, passes through a column, the stationary phase, that causes the metabolites to separate and elute at different times according to their solubility in both phases. Once the metabolites are eluted off the column, they are then ionized and sprayed into the mass spectrometer. Inside the mass spectrometer, metabolites are accelerated, and their mass to charge ratio is ultimately determined by their interaction with a magnetic field and detector. Subsequently, the metabolites hit a collision cell that breaks the parent metabolite into product metabolites. These product metabolites then interact with an additional magnetic field and detector to determine the products’ mass to charge (m/z) ratio. The output is a chromatogram that can be used to measure the amount of a specific analyte (Thomas et al. 2022). Technical improvements to the purification, separation, and detection of these compounds have been made over the last few decades (Aszyk et al. 2018; Famiglini et al. 2021). These improvements permit the analysis of these hormones in tissues that do not overaccumulate them, and with less chemical purification prior to analysis, thus allowing hormone profiling to occur within developmental and genetic studies in plants. For the first time, widely available chromatographic and mass spectroscopy technologies, paired with commercially available chemical standards, allow most laboratories with access to a triple-quadrupole mass spectrometer and high-performance liquid chromatography setup to quantify plant hormones.
In this review, we begin by describing the importance and benefits of studying hormone signaling in maize. We then briefly discuss historical hormone findings in maize, followed by a detailed description of historical and current methods for extracting and measuring maize hormones by chromatography and mass spectrometry. Finally, we discuss limitations to current techniques, as well as ways to optimize these techniques.
Why Maize?
Maize is one of the greatest contributors to calorie production in the world. Thus, direct applications of basic research findings in plant hormone regulation and their effects on plant growth can contribute to food security. Of particular interest is the production of shorter maize plants to increase plant densities, improve plant standing, reduce inputs, and increase harvest index in preparation for the effects of climate change. Quantification of hormone levels under different treatments and environments will allow for the development of new approaches to improve maize yields.
Maize has several advantages for studying hormone effects on growth and development. Of the model species with available genetic resources and annual generation times, maize is a relatively large plant and accumulates substantial tissue for hormone extraction. There is a large and collaborative maize research community that manages multiple seed repositories with natural variants and mutants (http://maizecoop.cropsci.uiuc.edu/; https://www.ars-grin.gov/; Settles et al. 2007; Lu et al. 2018; Marcon et al. 2020). Among these mutants are several with known blocks in the production of ABA, auxin, BR, GA, JA, SA, and strigolactone. There are also mutants with defects in ABA, auxin, BR, CK, ethylene, GA, and JA signaling pathways. These mutant lines present an opportunity to learn how various hormones interact to influence the accumulation of each other. For example, we found that maize mutants defective in the production of either BR or GA require the synthesis of the other hormone to fully exhibit mutant phenotypes (Best et al. 2016, 2017). In double mutant and inhibitor experiments, epistasis was identified between mutant genes in these two pathways. For example, if you block GA synthesis, some of the phenotypes observed in BR-deficient mutants were suppressed. Likewise, if you block BR biosynthesis, some of the defects observed in GA-deficient mutants were suppressed. Unfortunately, these studies failed to measure hormone levels, and as a result, it is unknown whether GA and BR influence each other's synthesis or merely gate each other's responses, by altering the plant's sensitivity to the relevant hormone. The large size of the maize plant body may also help in the identification of additional hypothesized, but as yet unidentified, potential plant hormones that have been defined genetically. Examples of these hypothesized hormones include the signals acted on by the CYP78A family of P450 enzymes (Anastasiou et al. 2007), a signal with apparent root-to-shoot communication defined by the bypass1 mutant (Van Norman et al. 2004), and the endogenous ligand for the KAI2 receptor (Villaécija-Aguilar et al. 2019), none of which have chemical identities as of this writing.
The foundational understanding of hormones is derived from physiological and biochemical studies. Recent advances in molecular genetics have led to hormone-related studies based on genetic homology of genes and mutant phenotypes. In many cases, including our own work, hormone effects in maize have been inferred from the phenotypes of mutants in genes homologous to known hormone biosynthetic enzymes or signal transduction components, without a coincident measurement of hormone levels. Such studies can cloud understanding and create controversy, particularly as feedback between the perception and biosynthesis of signaling compounds is known for multiple pathways. For example, a tall plant may be tall because of a loss of GA catabolism (e.g., a GA2 oxidase mutant), increased GA production (e.g., overexpression of ent-kaurene oxidase), or loss of a negative regulator of GA signaling (e.g., a DELLA domain transcription factor mutant), making it difficult to draw conclusions based on genetics alone. In the example above, the first two mutants would be expected to accumulate GA, whereas the last mutant would be expected to lack GA. Knowing the hormone levels in specific mutant tissues allows a more in-depth understanding of the mutation. Additionally, the biosynthetic pathways of hormones are interlinked through feedback and signaling interactions that cannot be fully explained by transcriptomics or proteomics. Therefore, measuring a single hormone is insufficient to test a hypothesis successfully, and a method to simultaneously measure as many hormone metabolites as possible allows for a more accurate interpretation of experimental findings. The capability to measure multiple hormones in a single extraction allows us to assess the levels of multiple hormones in parallel and enables researchers to study the interplay between these hormones and their signaling during development.
Extraction/Measurement of Hormones
Much of our understanding of hormone structure and accumulation is derived from the study of compounds extracted using a solvent that efficiently recovers a single hormone class, which are then analyzed by gas chromatography (GC)/mass spectrometry (MS). In many cases, this analysis required derivatization, or chemical modification, of these small molecules. Derivatization can aid in chromatographic separation and detection. It can also improve the ionization of compounds during MS, resulting in consistent and accurate detection of metabolite classes. During MS, compounds are ionized with the application of a negative or positive charge. Some compounds will readily ionize as either a positive or negatively charged molecule. Derivatization is particularly valuable for compounds that are accumulated at low concentrations or that ionize poorly in mass spectral analysis. Derivatization with a moiety that carries a charge can solve the problem of poor ionization. Derivatization does require more steps in the experimental procedure and the chemical reaction that is used to modify the compound typically only affects a subset of molecules in a sample. If the chemical reactions are sufficiently specialized, detection of the modifications can be the strongest signal available.
LC/MS has improved greatly in the last three decades in terms of chromatographic reproducibility, MS sensitivity, and mass accuracy of the detector. These advances have shifted attention from GC to LC for hormone analysis. This shift has not only changed the mobile phase for chromatography but has also removed the necessity of derivatization for analysis of most hormones. LC allows the measurement of endogenous molecules and the ability to simultaneously detect multiple metabolites in the same sample. For metabolites to be analyzed by mass spectrometry, they need to ionize efficiently. Many plant hormones are negatively charged or weakly polar and efficiently ionized as negatively charged ions. The one notable exception are the BRs, which lack a charged moiety, and therefore require derivatization. Adaptation of a method for the detection of sterols, which are intermediates in the hormone synthesis pathway, and steroid hormones from humans (Honda et al. 2010) has permitted us to detect BRs as picolinate esters (Dilkes and Best 2025b, in this collection). This derivatization, which is specific to sterol compounds with a 3′ hydroxyl moiety, was shown to both improve ionization and permit quantitation of BRs (Yang et al. 2018; Tang et al. 2022), and is described in our associated protocol (Dilkes and Best 2025a). Other approaches for derivatizing BRs at the 3′ hydroxyl that are compatible with LC/MS have also been used, including 3-(dimethylamino)-phenylboronic acid (DMAPBA) derivatization (Xin et al. 2013).
Standards Limitation—Chemical Synthesis Gap
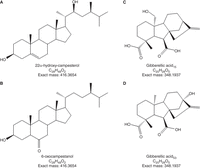
LC/MS experiments detect “mass features,” which are the result of ionized molecules hitting a detector within a time window of the chromatography. As such, there may be many mass features with the same chemical formula as a compound of interest, and a feature with the correct mass of a hormone is insufficient evidence that this feature is due to the observation of that hormone. Comparison of both the MS spectra and chromatographic retention time of a feature to a known standard is typically required for compound identification. The difficulty and expense of producing chemically known standards has limited the number of laboratories and teams that can study plant regulatory molecules and hormones. A “chemical synthesis gap,” where proposed chemicals have not yet been or cannot readily be synthesized, can occur when there is purification and detection of a molecule that does not have a synthesized standard to which it can be compared for chemical identification during MS analyses. For some hormone classes, particularly the BRs and GAs, multiple compounds have the same mass, because they consist of the same chemical formula arranged in a different structure. Thus, differences in retention time caused by differences in structure (e.g., different orientation of an –OH group resulting in a change in interaction with the column and mobile phase) is key to determining molecule identity (see Fig. 1) and, unlike m/z, which can be arrived at theoretically, retention times on a column must be measured using bona fide standards. In addition to the traditional use of known chemical standards, metabolites purified from mutant plants with disruptions in hormone biosynthetic pathways can be used to identify hormones. Tracking the accumulated and missing metabolites from mutant and wild-type tissues can further help in the identification of peaks at novel retention times.
Chemical structure of pairs of brassinosteroid and gibberellin metabolites with the same mass and chemical formula but different chemical structures. Chemical structure of 22α-hydroxy-campesterol (A), 6-oxocampestanol (B), gibberellic acid15 (C), and gibberellic acid53 (D).
The addition of deuterium-labeled standards can further improve LC/MS analysis and allows for absolute quantification of the hormones. Deuterated compounds are added to ground tissue at known concentrations and are nearly chemically identical to nondeuterated hormones during extraction and separation. They can be distinguished by mass during analysis and used to create a standard curve. In the absence of deuterium-labeled standards for every individual metabolite in a hormone class (e.g., in the presence of a chemical synthesis gap), indirect quantification of hormones can be carried out using a standard curve from a chemically similar compound. For example, our associated protocol (Dilkes and Best 2025b) uses a deuterated GA53 (GAs are numbered in order of discovery) standard to produce a standard curve for the entire GA class of metabolites. Using a representative metabolite as a standard is an effective approach, especially when the cost and availability of deuterated bona fide standards are prohibitive. This approach allows quantification of one chemical hormone to act as the proxy for others in the same class, and thus the accumulation of a whole class of molecules can be assessed. A potential pitfall in the approach that can confound interpretations is that even though compounds are in the same hormone class and are structurally similar, they may have altered ionization or differential extraction from the compound used for the standard curve.
Purification and Signal Improvement
The total amount of hormone that can be loaded onto a column and sprayed into a mass spectrometer limits the amount of signal that can be observed. Prior methods required purification of each hormone away from other metabolites before detection. Improvements in the sensitivity and mass accuracy of instruments to detect these hormones now allow for simultaneous observance of multiple hormones in the same sample. However, this less targeted approach can result in unwanted metabolites that can interfere with the detection and quantification of the wanted hormones remaining in the analyzed sample. The addition of a solid phase extraction step improves the signal to noise ratio by removing some unwanted metabolites from the samples. Our method uses graphitized carbon black to achieve this purification, but other sorbents could be used, such as activated carbons. Carbon black can react with hormones and, if left in contact with the sample for too long, will reduce signal levels.
Derivatization is another form of signal improvement. We propose that the combination of derivatization, targeted (tandem MS/MS) and untargeted (quadrupole time of flight [QTOF] MS) profiling, and metabolite measurements in mutants can ultimately identify all hormone pathway intermediates. The QTOF MS allows detection of all metabolites in a sample without previously knowing their exact masses prior to analysis. In targeted MS/MS analyses, extraction and separation of specific metabolites are carried out and when metabolites of the expected mass are detected, these metabolites are further fragmented, and their fragments are then detected to provide further structural detail. All forms of extraction and derivatization are designed to bias the signal for a specific hormone class. However, extraction and derivatization can also cause some distortion of that signal, which can be problematic when looking at multiple metabolites within the same class. For example, the method we present in the associated BR protocol (Dilkes and Best 2025a) uses derivatization at the 3′ hydroxyl of sterols with picolinic acid. Some BRs have a 3′ ketone group at this location and therefore will not derivatize and will not be detected in this protocol. There are alternate methods to derivatize these 3′ keto sterols, such as the Amplifex Keto Reagent, which derivatizes 3′ keto steroids with a permanently changed quaternary amine that stabilizes the structure and improves ionization in positive mode (Zhu et al. 2015). This derivatization allows the efficient ionization of the few BR intermediates with a 3′ ketone, including the pregnane derivatives formed by side-chain cleavage of the steroid by enzymes in the plant (Yang et al. 2018; Carroll et al. 2023; Kunert et al. 2023). If desired, a second round of analysis incorporating this method will allow for a global analysis inclusive of the 3′ keto steroids (Dilkes and Best 2025b, in this collection).
These methods are not exclusively useful for previously identified compounds. For example, performing an untargeted LC/MS profiling of the picolinate derivatives of all steroids, and not just the known BRs, can allow a search of the data for the picolinate fragment as an informatic anchor, and the potential identification of novel sterols. This technique was previously successful on tissue from the Setaria BR-deficient cytochrome P450 724 (CYP724) mutant bristleless1, which accumulated a C21 pregnane (Yang et al. 2018). In underivatized samples, we also detected a mass consistent with the 3′ ketone derivative of this same compound and hypothesized the existence of a plant desmolase (Yang et al. 2018). Recently, the pathway to produce these compounds was elucidated (Carroll et al. 2023; Kunert et al. 2023), confirming the hypothesized pathway uncovered by untargeted metabolite profiling of a hormone biosynthetic mutant.
Many more such opportunities for characterizing and identifying hormones exist across the plant kingdom. For example, the ultimate bioactive products of the BR biosynthetic pathway in maize and other monocots are still unclear. The diversification of the cytochrome P450 90s (CYP90) genes that encode the BR biosynthetic pathway enzymes that we are familiar with in flowering plants occurred after the split between vascular and nonvascular plants. As a result, the CYP90s in plants other than the angiosperms may synthesize different polyhydroxylated sterols that act as BRs in these plants. GAs may also be different across plant taxa. For example, the sex-determining antheridogens in some homosporous ferns are GAs. To date, all chemically identified antheridogens are GAs, but many antheridogens are not yet chemically identified and may be novel GAs or other signaling molecules (Takeno 1991; Tanaka et al. 2014). Additional opportunities for chemical identification include the metabolic targets of the CYP78A enzymes, the KLUH pathway, members of which are also as yet unidentified (Anastasiou et al. 2007; Stransfeld et al. 2010). Studying mutants in these pathways in maize, where greater volumes of tissue are readily obtained for metabolite purification than in other plants, may aid in the chemical identification of this and other proposed hormones.
CONCLUSION
Using maize as a model system presents the perfect combination of large vegetative plant size, available genetic resources, and an inclusive scientific community to carry out hormone studies in plants. Together with the wide availability of LC/MS instrumentation, the sensitivity gains afforded by recent technical developments should allow any researcher to incorporate measurements of these previously difficult to quantify phytohormones into their research toolbox. We hope these techniques usher in a new era of understanding the roles of hormones in plant growth and development.
COMPETING INTEREST STATEMENT
The authors declare no competing interests. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture. The U.S. Department of Agriculture is an equal opportunity provider and employer.
AUTHOR CONTRIBUTIONS
Conceptualization: B.P.D. and N.B.B. Investigation: B.P.D. and N.B.B. Writing—original draft: B.P.D. and N.B.B. Writing—review and editing: B.P.D. and N.B.B.
Acknowledgments
We thank Amber Hopf Jannasch, Bruce Cooper, and Jim Elder for helping in setting up this protocol.
Footnotes
-
From the Maize collection, edited by Candice N. Hirsch and Marna D. Yandeau-Nelson. The entire Maize collection is available online at Cold Spring Harbor Protocols and can be accessed at https://cshprotocols.cshlp.org/.
REFERENCES
*Reference is also in this subject collection.